Introduction:
A protein domain is a portion of a protein sequence that has been relatively conserved through evolution. Multiple protein domains may be found on one polypeptide [1]. Domain analysis is considered to assist gene ontology in classifying perforin- 1 and identifying related biological processes.
Results:
Discussion:
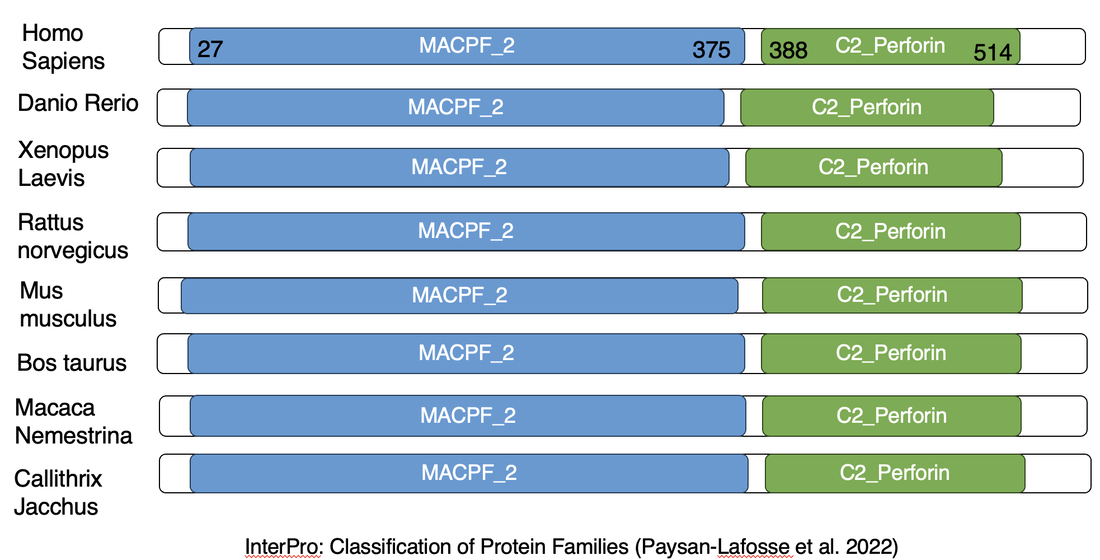
One major protein domain in perforin is the membrane attack complex/perforin. This is the domain responsible for pore formation as described on the gene ontology page [2]. The calcium domain in perforin has no identified localization point, but it is involved in leukocyte mediated cytotoxicity similar to that of the membrane attack perforin domain. Evidence suggests perforin requires calcium to bind to the cell membrane, so this domain assists in this process [2]. Running the FASTA sequence information for identified homologs through Interpro scan provided results suggesting that the homologs have similar pore formation and binding to cellular membranes across invertebrates, and that this biological process is well conserved.
One major protein domain in perforin is the membrane attack complex/perforin. This is the domain responsible for pore formation as described on the gene ontology page [2]. The calcium domain in perforin has no identified localization point, but it is involved in leukocyte mediated cytotoxicity similar to that of the membrane attack perforin domain. Evidence suggests perforin requires calcium to bind to the cell membrane, so this domain assists in this process [2]. Running the FASTA sequence information for identified homologs through Interpro scan provided results suggesting that the homologs have similar pore formation and binding to cellular membranes across invertebrates, and that this biological process is well conserved.
References:
[1] Wang, Yan, et al. “Protein Domain Identification Methods and Online Resources.” Computational and Structural Biotechnology Journal, U.S. National Library of Medicine, 2 Feb. 2021, www.ncbi.nlm.nih.gov/pmc/articles/PMC7895673/.
[2] Paysan-Lafosse T, Blum M, Chuguransky S, Grego T, Pinto BL, Salazar GA, Bileschi ML, Bork P, Bridge A, Colwell L, Gough J, Haft DH, Letunić I, Marchler-Bauer A, Mi H, Natale DA, Orengo CA, Pandurangan AP, Rivoire C, Sigrist CJA, Sillitoe I, Thanki N, Thomas PD, Tosatto SCE, Wu CH, Bateman A. InterPro in 2022. Nucleic Acids Research, Nov 2022, (doi: 10.1093/nar/gkac993)
[1] Wang, Yan, et al. “Protein Domain Identification Methods and Online Resources.” Computational and Structural Biotechnology Journal, U.S. National Library of Medicine, 2 Feb. 2021, www.ncbi.nlm.nih.gov/pmc/articles/PMC7895673/.
[2] Paysan-Lafosse T, Blum M, Chuguransky S, Grego T, Pinto BL, Salazar GA, Bileschi ML, Bork P, Bridge A, Colwell L, Gough J, Haft DH, Letunić I, Marchler-Bauer A, Mi H, Natale DA, Orengo CA, Pandurangan AP, Rivoire C, Sigrist CJA, Sillitoe I, Thanki N, Thomas PD, Tosatto SCE, Wu CH, Bateman A. InterPro in 2022. Nucleic Acids Research, Nov 2022, (doi: 10.1093/nar/gkac993)