Introduction to Transcriptomics:
Transcriptomics is the study of all RNA extracts in a system. RNA reveals regulatory processes in the genome as well as can elucidate previously unidentified gene functions. Methods for transcriptomics includes RNA- seq, which involves sequencing RNA transcripts from cDNA. Another method, microarrays, involves multiple short nucleotide reads/probes that flouresce at different levels of intensity, which is proportional to the abundance of that read [1].
Transcriptomics is the study of all RNA extracts in a system. RNA reveals regulatory processes in the genome as well as can elucidate previously unidentified gene functions. Methods for transcriptomics includes RNA- seq, which involves sequencing RNA transcripts from cDNA. Another method, microarrays, involves multiple short nucleotide reads/probes that flouresce at different levels of intensity, which is proportional to the abundance of that read [1].
Discussion:
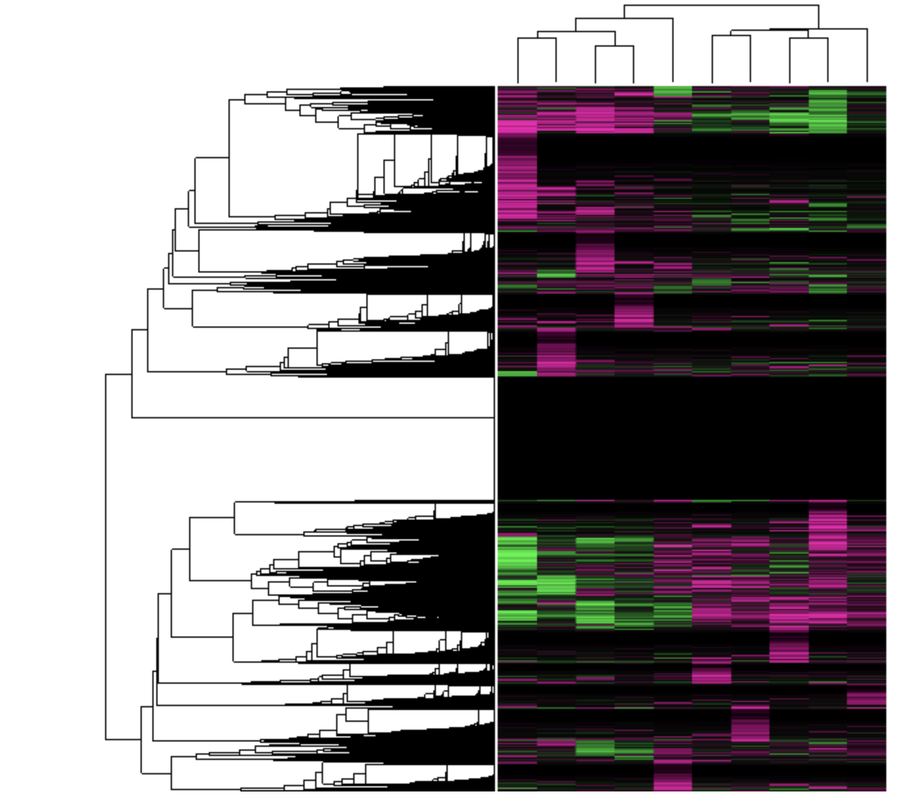
Transcriptomic methods were used in a similar study (see above cluster analysis). Although the study focuses on analysis of juvenile idiopathic arthritis (JIA), fHLH patients were brought into sampling due to similar NK overactivation and increased serum ferritin concentrations. Microarray was performed on patient samples, and cluster analysis was completed. Clustering of probes identified with a large production of red blood cells supported the hypothesis that fHLH and sJIA have similar pathways associated [2]. To determine the relationship between ferritin levels and fHLH, a similar method is proposed to cluster probes associated with iron homeostasis and perforin-1 formation after microarray.
Transcriptomic methods were used in a similar study (see above cluster analysis). Although the study focuses on analysis of juvenile idiopathic arthritis (JIA), fHLH patients were brought into sampling due to similar NK overactivation and increased serum ferritin concentrations. Microarray was performed on patient samples, and cluster analysis was completed. Clustering of probes identified with a large production of red blood cells supported the hypothesis that fHLH and sJIA have similar pathways associated [2]. To determine the relationship between ferritin levels and fHLH, a similar method is proposed to cluster probes associated with iron homeostasis and perforin-1 formation after microarray.
References:
[1] Lowe, Rohan, et al. “Transcriptomics Technologies.” PLoS Computational Biology, U.S. National Library of Medicine, 18 May 2017, www.ncbi.nlm.nih.gov/pmc/articles/PMC5436640/.
[2] Hinze, Claas H, et al. “Immature Cell Populations and an Erythropoiesis Gene-Expression Signature in Systemic Juvenile Idiopathic Arthritis: Implications for Pathogenesis.” Arthritis Research & Therapy, U.S. National Library of Medicine, 2010, www.ncbi.nlm.nih.gov/pmc/articles/PMC2911917/.
Image used. “GDS Cluster Analysis.” National Center for Biotechnology Information, U.S. National Library of Medicine, www.ncbi.nlm.nih.gov/geo/gds/analyze/analyze.cgi?ID=GDS4142. Accessed 9 Apr. 2024.
[1] Lowe, Rohan, et al. “Transcriptomics Technologies.” PLoS Computational Biology, U.S. National Library of Medicine, 18 May 2017, www.ncbi.nlm.nih.gov/pmc/articles/PMC5436640/.
[2] Hinze, Claas H, et al. “Immature Cell Populations and an Erythropoiesis Gene-Expression Signature in Systemic Juvenile Idiopathic Arthritis: Implications for Pathogenesis.” Arthritis Research & Therapy, U.S. National Library of Medicine, 2010, www.ncbi.nlm.nih.gov/pmc/articles/PMC2911917/.
Image used. “GDS Cluster Analysis.” National Center for Biotechnology Information, U.S. National Library of Medicine, www.ncbi.nlm.nih.gov/geo/gds/analyze/analyze.cgi?ID=GDS4142. Accessed 9 Apr. 2024.